Cellular mechanisms of copper transport and delivery to cytochrome oxidase
Copper is an essential micronutrient and functions as cofactor in many important enzymes like Cytochrome c oxidases or the human dopamine beta-hydroxylase. Despite the undeniable necessity for copper uptake, free copper is highly toxic and all cells use sophisticated transport and storage systems for maintaining Cu homeostasis and detoxification. Mutations in these systems are associated with severe genetic diseases in humans like Morbus Wilson or Menkes disease.
One of the most important copper-dependent proteins is the membrane-embedded cytochrome c oxidase (Cox), which terminates the respiratory chain in aerobic organisms. Here, Cu is part of the active centre of Cox and required for the reduction of O2. Although the presence of Cu in the active centre of Cox is known since decades, the exact mechanisms by which Cu is inserted into the active centre is still not known. For analysing this, we are using the cbb3-type cytochrome c oxidase of the α-proteobacterium Rhodobacter capsulatus as a model enzyme. This cytochrome oxidase is not only a perfect model system; it is also of high medical importance because this type of cytochrome oxidase is present in many pathogenic bacteria like Neisseria meningitides, Campylobacter jejuni or Helicobacter pylori.
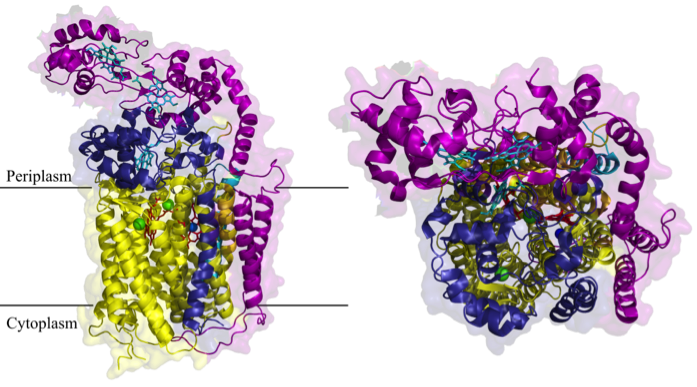
Fig.1 Structure of cbb3-type cytochrome c oxidase of Pseudomonas stutzeri (Buschmann et al., 2012)
In recent studies we have identified several proteins that are involved in copper homeostasis and delivery to cbb3 Cox (Table 1).
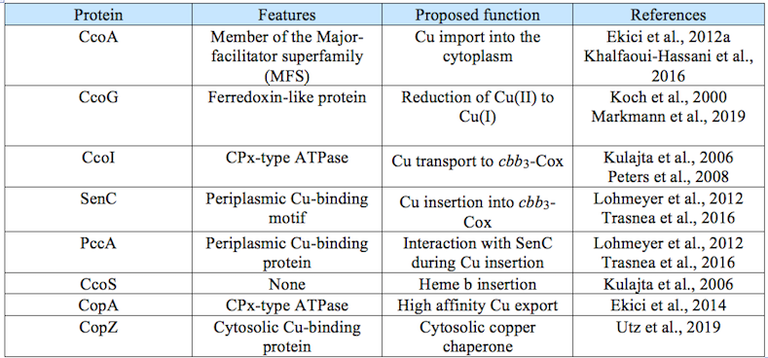
Table 1: Proteins involved in Cu homeostasis in Rhodobacter capsulatus.
Based on the biochemical characterizations of these proteins, a tentative model of Cu transport in R. capsulatus was proposed (Fig. 2): Copper is imported as Cu(II) via the novel MFS-like protein CcoA and subsequently reduced to Cu(I) by the ferredoxin-like protein CcoG. Cu(I) is either bound by the cytosolic chaperone CopZ or exported back to the periplasm by the P1B-type ATPase CcoI. In the periplasm, the coordinated activity of two Cu(I)-specific periplasmic chaperones, PccA and SenC, probably insert Cu into the catalytic subunit CcoN of cbb3-type Cox. Excess of Cu is exported to the periplasm by another P1B-type ATPase, CopA, which is not required for cbb3-type Cox assembly, but for preventing Cu toxicity.
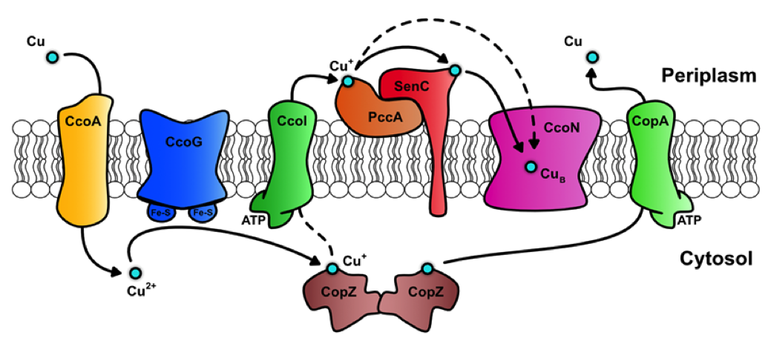
Fig. 2 Copper homeostasis in Rhodobacter capsulatus
Even though our understanding of the Cu transport pathways in R. capsulatus has dramatically improved during the last years, the molecular details of this intricate and complex protein interaction network that finally sustain the successful assembly of cbb3 Cox remain to be clarified. This interdisciplinary project aims to delineate copper trafficking to cytochrome oxidase and to define the precise functions of the identified copper binding proteins by biochemical, biophysical, cell biological and genetic methods.
References
[1] Lohmeyer E, Schröder S, Pawlik G, Trasnea PI, Peters A, Daldal F, Koch H-G (2012) The ScoI homologue SenC is a copper binding protein that interacts directly with the cbb3-type cytochrome oxidase in Rhodobacter capsulatus. Biochim Biophys Acta 1817:2005-2007.
[2] Ekici S, Turkarslan S, Pawlik G, Dancis A, Baliga NS, Koch H-G, Daldal F (2014) Intracytoplasmic Copper Homeostasis Controls Cytochrome c Oxidase Production. mBio 5:e01055-13.
[3] Trasnea, P.I., Utz, M., Khalfaoui-Hassani, B., Lagies, S., Daldal, F. and Koch, H.G. (2016) Cooperation between two periplasmic copper chaperones is required for full activity of the cbb3-type cytochrome c oxidase and copper homeostasis in Rhodobacter capsulatus. Mol. Microbiology 100, 345-361.
Publications
Denks, K, Sliwinski, N, Erichsen, V, Borodkina, B, Origi, A and Koch, HG (2017) The signal recognition particle contacts uL23 and scans substrate translation inside the ribosomal tunnel. Nature Microbiology, 2, 16265.
Khalfaoui-Hassani, B., Wu, H., Blaby-Haas, C., Zhang, Y., Sandri, F., Verissimo, A.F., Koch, H.-G., and Daldal, F. (2018). Widespread Distribution and Functional Specificity of the Copper Importer CcoA: Distinct Cu Uptake Routes for Bacterial Cytochrome c Oxidases. mBIO 9, e00065-18.
Trasnea, P.I., Andrei, A., Markmann, D., Utz, M., Khalfaoui-Hassani, B.; Selamoglu, N., Daldal, F., and Koch, H.G. (2018). A copper relay system involving two periplasmic chaperones drives cbb3 type cytochrome c oxidase biogenesis in Rhodobacter capsulatus. ACS Chemical Biology 13, 1388-1397.
Utz, M. Andrei, A., Milanov, M., Trasnea, PI, Marckmann, D., Daldal, F., Koch, H.-G. (2019) The Cu chaperone CopZ is required for Cu homeostasis in Rhodobacter capsulatus and influences cytochrome cbb3 oxidase assembly. Mol Microbiology 111(3):764-783.
Zhang, Y., Blaby-Haas, C.E., Steimle, S., Verissimo, A.F., Garcia-Angulo, V., Koch, H.G., Daldal, F., Khalfaoui-Hassani, B (2019) Cu Transport by the Extended Family of CcoA-like Transporters (CalT) in Proteobacteria. Scientific reports 9:1208.
Marckmann, D., Trasnea, P.I, Schimpf, J., Winterstein, C., Andrei, A., Schmollinger, S., Blaby-Haas, C.E., Friedrich, T., Daldal, F., Koch, H.-G. (2019) The cbb3-type cytochrome oxidase assembly factor CcoG is a widely distributed cupric reductase. Proc. Natl. Acad. Sci. USA, 116, 21166-21175.
Selamoglu, N., Önder, Ö., Öztürk, Y., Khalfaoui-Hassani, B., Blaby-Haas, C.B., Garcia, B., Koch, H.G. and Daldal, F. (2020) Comparative Differential Cuproproteomes of Rhodobacter capsulatus Reveal Novel Copper Homeostasis Related Proteins. Metallomics in press.
Contact
Prof. Dr. Hans-Georg Koch
Institute for Biochemistry and Molecular Biology
Stefan-Meier-Str. 17
79104 Freiburg
Phone: +49 (761) 203 5250
Fax: +49 (761) 203 5289
hans-georg.koch@biochemie.uni-freiburg.de
http://www.biochemie.uni-freiburg.de/ag-en/koch/koch/



